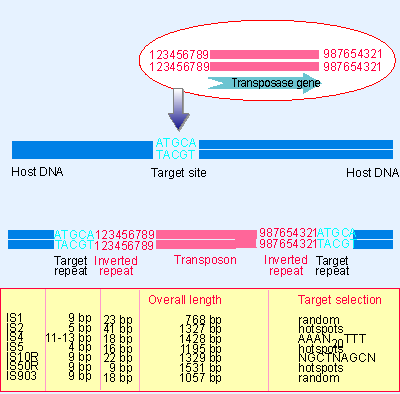
Figure 1 from Structure-function analysis of the inverted terminal repeats of the sleeping beauty transposon. | Semantic Scholar
PLOS ONE: detectIR: A Novel Program for Detecting Perfect and Imperfect Inverted Repeats Using Complex Numbers and Vector Calculation
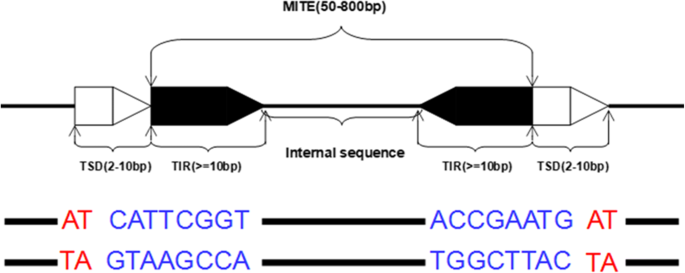
MiteFinderII: a novel tool to identify miniature inverted-repeat transposable elements hidden in eukaryotic genomes | BMC Medical Genomics | Full Text

Secondary structures formed by inverted repeats and AT- and CG-rich... | Download Scientific Diagram
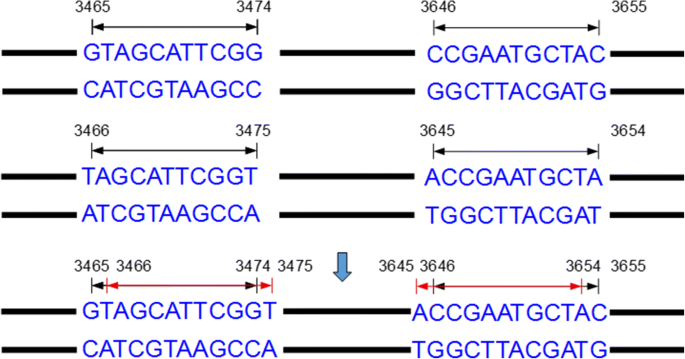
MiteFinderII: a novel tool to identify miniature inverted-repeat transposable elements hidden in eukaryotic genomes | BMC Medical Genomics | Full Text
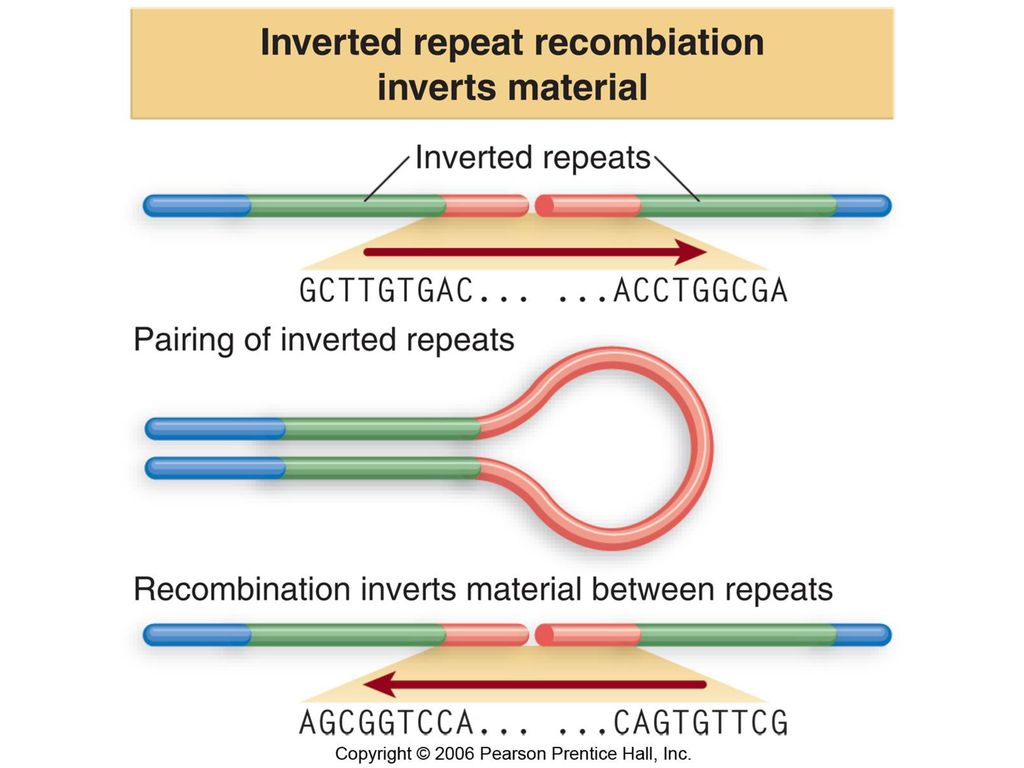
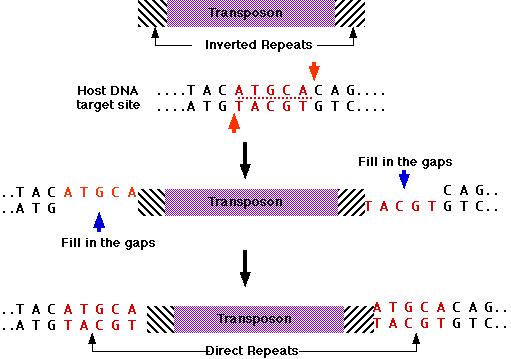
Figure: Title: Transposons have inverted repeats and generate target repeats Caption: Transposons have inverted terminal repeats and generate direct. - ppt download

Multiple Promoter Inversions Generate Surface Antigenic Variation in Mycoplasma penetrans | Journal of Bacteriology

Recognition of DNA by Fur: a Reinterpretation of the Fur Box Consensus Sequence | Journal of Bacteriology